Boosting - GBM
참고
앙상블예제
GBM(Gradient Boosting Machine)
부스팅 앙상블 (Boosting Ensemble) 2-1: Gradient Boosting for Regression
부스팅이란?
여러 개의 약한 학습기(weak learner)를 순차적으로 학습 → 예측을 하면서 잘못 예측한 데이터에 가중치를 부여해서 오류를 개선하면서 나아가는 방식
즉, 이전 학습기의 잔차 (resuidual)를 보완하는 방식이다.
GBM(Gradient Boosting Machine) 이란
가중치를 업데이트 할때 경사 하강법(Gradient Descent)을 이용하는 알고리즘이다.
- 경사하강법(Gradient Descent) : 손실 함수의 최솟값을 찾는 최적화(optimization) 방법 중 하나
기본 개념은 함수의 기울기(경사)를 구하고 경사의 반대 방향으로 계속 이동시켜 극값
에 이를 때까지 반복시키는 것이다. → 머신러닝에서 예측값과 정답값 간의 차이인 손실함수의 크기를 최소화시키는 파라미터를 찾기 위해 사용한다. 수식을 통해 손실 함수의 값이 거의 변하지 않을 때까지 가중치를 업데이트하는 과정을 반복한다. 이처럼 손실 함수 그래프에서 값이 가장 낮은 지점으로(=손실 함수의 최솟값) 경사를 타고 하강하는 기법을 경사 하강법이라고 한다.
수식을 통해 손실 함수의 값이 거의 변하지 않을 때까지 가중치를 업데이트하는 과정을 반복한다. 이처럼 손실 함수 그래프에서 값이 가장 낮은 지점으로(=손실 함수의 최솟값) 경사를 타고 하강하는 기법을 경사 하강법이라고 한다.
GBM 의 알고리즘 방식
GBM은 약한 학습기를 잔차 자체에 적합하고, 이전의 예측값에 예측한 잔차를 더해 주어 예측값을 새로이 업데이트하는 방식이다.
- 간단한 모델 A를 통해 y를 예측
- 예측하고 남은 잔차 (residual)을 다시 B라는 모델을 통해 예측
- A+B 모델을 통해 y를 예측
→ 잔차는 계속해서 줄어들게되고, training set을 잘 설명하는 예측 모형을 만들 수 있게 된다.
-
GBM 은 약한 학습기로 restricted tree 를 사용한다.
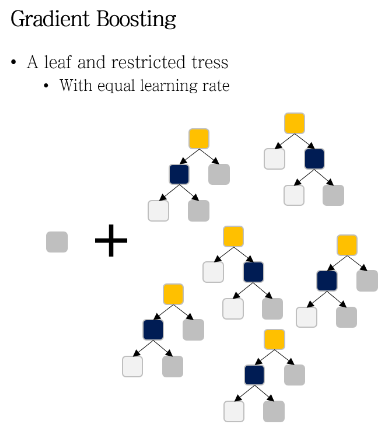
restricted tree란, maximum number of leaves로 성장에 제한을 둔 decision tree이다.
즉 GBM 은 작은 decision tree 들로 이루어져 있다.
장점 : 성능이 좋다
단점 : 과대적합할 가능성이 높고, 연산이 많아 시간이 오래 걸린다.
실습해보기
- Pima Indians Dataset 이용
- 전처리
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
df = pd.read_csv("http://bit.ly/data-diabetes-csv")
print(df.shape)
# Insulin 결측치 제거
df.loc[df["Insulin"] > 0 , "Insulin_fill"] = df["Insulin"]
median_sr = df.groupby("Outcome")["Insulin_fill"].median()
print(median_sr[0],median_sr[1])
df.loc[(df["Insulin_fill"].isna()) & (df["Outcome"] == 0), "Insulin_fill"] = median_sr[0]
df.loc[(df["Insulin_fill"].isna()) & (df["Outcome"] == 1), "Insulin_fill"] = median_sr[1]
# train , test 데이터 셋 만들기
label_name = "Outcome"
feature_names = df.columns.tolist()
feature_names.remove(label_name)
feature_names.remove("Insulin")
X = df[feature_names]
y = df[label_name]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42, stratify=y, test_size=0.2)- GBM 실습
import time # 시간측정을 위해 import
from sklearn.ensemble import GradientBoostingClassifier
# 모델생성
gbm = GradientBoostingClassifier(random_state=42)
start_gbm = time.time() # 시작시간
gbm.fit(X_train, y_train) # training
gbm_predict = gbm.predict(X_test) # 예측
end_gbm = time.time() # 끝낸시간
gbm_accuracy =(y_test == gbm_predict).mean() # 예측결과
print(" GradientBoosting 걸린시간 : ", end_gbm - start_gbm , " 예측점수 : ", gbm_accuracy)
>>> GradientBoosting 걸린시간 : 0.2343740463256836 예측점수 : 0.8766233766233766- DecisionTree 와 비교
from sklearn.tree import DecisionTreeClassifier
# 모델생성
model = DecisionTreeClassifier(random_state=42)
start_d = time.time() # 시작시간
model.fit(X_train, y_train) # training
y_predict = model.predict(X_test) # 예측
end_d = time.time() # 끝낸시간
accuracy = (y_test == y_predict).mean() # 예측결과
print(" DecisionTree 걸린시간 : ", end_d - start_d , " 예측점수 : ", accuracy)
>>> DecisionTree 걸린시간 : 0.00997304916381836 예측점수 : 0.8571428571428571GradientBoosting 모델이 성능이 높은대신 시간이 오래걸리는 것을 볼 수 있다!
파라메터
# GBM 의 파라메터 Default
GradientBoostingClassifier(
*,
loss='deviance',
learning_rate=0.1,
n_estimators=100,
subsample=1.0,
criterion='friedman_mse',
min_samples_split=2,
min_samples_leaf=1,
min_weight_fraction_leaf=0.0,
max_depth=3,
min_impurity_decrease=0.0,
init=None,
random_state=None,
max_features=None,
verbose=0,
max_leaf_nodes=None,
warm_start=False,
validation_fraction=0.1,
n_iter_no_change=None,
tol=0.0001,
ccp_alpha=0.0,
)-
GBM 은 내부에 DecisionTree 가 있기 때문에, DecisionTree가 가지고 있는 파라메터의 속성을 가지고 있다.
- max_features
- max_depth
- min_sample_leaf
- min_sample_split
-
GBM 파라메터 설명
- loss : gradient descent 에서 사용할 손실함수. default 는 ‘deviance’
- learing_rate : 학습률. 실수값을 가진다. default 는 0.1 이다. 값이 너무 적으면 학습이 더디고 너무 크면 과대적합이 일어날 수 있다. 보통 0.05~0.2 사이의 값을 가진다.
- subsample : weak learner 가 학습에 사용하는 데이터의 샘플링 비율이다. default 값은 1 이고 이는 전체 데이터를 기반으로 한다는 뜻이다. 0.7 이면 70%를 사용한다는 뜻이다.
-
파라메터 자세히 보기
Init signature:
GradientBoostingClassifier(*, loss='deviance', learning_rate=0.1, n_estimators=100, subsample=1.0, criterion='friedman_mse', min_samples_split=2, min_samples_leaf=1, min_weight_fraction_leaf=0.0, max_depth=3, min_impurity_decrease=0.0, init=None, random_state=None, max_features=None, verbose=0, max_leaf_nodes=None, warm_start=False, validation_fraction=0.1, n_iter_no_change=None, tol=0.0001, ccp_alpha=0.0,)Docstring:
Gradient Boosting for classification.
GB builds an additive model in a
forward stage-wise fashion; it allows for the optimization of
arbitrary differentiable loss functions. In each stage ``n_classes_``
regression trees are fit on the negative gradient of the
binomial or multinomial deviance loss function. Binary classification
is a special case where only a single regression tree is induced.
Read more in the :ref:`User Guide <gradient_boosting>`.
Parameters
----------
loss : {'deviance', 'exponential'}, default='deviance'
The loss function to be optimized. 'deviance' refers to
deviance (= logistic regression) for classification
with probabilistic outputs. For loss 'exponential' gradient
boosting recovers the AdaBoost algorithm.
learning_rate : float, default=0.1
Learning rate shrinks the contribution of each tree by `learning_rate`.
There is a trade-off between learning_rate and n_estimators.
n_estimators : int, default=100
The number of boosting stages to perform. Gradient boosting
is fairly robust to over-fitting so a large number usually
results in better performance.
subsample : float, default=1.0
The fraction of samples to be used for fitting the individual base
learners. If smaller than 1.0 this results in Stochastic Gradient
Boosting. `subsample` interacts with the parameter `n_estimators`.
Choosing `subsample < 1.0` leads to a reduction of variance
and an increase in bias.
criterion : {'friedman_mse', 'squared_error', 'mse', 'mae'}, default='friedman_mse'
The function to measure the quality of a split. Supported criteria
are 'friedman_mse' for the mean squared error with improvement
score by Friedman, 'squared_error' for mean squared error, and 'mae'
for the mean absolute error. The default value of 'friedman_mse' is
generally the best as it can provide a better approximation in some
cases.
.. versionadded:: 0.18
.. deprecated:: 0.24
`criterion='mae'` is deprecated and will be removed in version
1.1 (renaming of 0.26). Use `criterion='friedman_mse'` or
`'squared_error'` instead, as trees should use a squared error
criterion in Gradient Boosting.
.. deprecated:: 1.0
Criterion 'mse' was deprecated in v1.0 and will be removed in
version 1.2. Use `criterion='squared_error'` which is equivalent.
min_samples_split : int or float, default=2
The minimum number of samples required to split an internal node:
- If int, then consider `min_samples_split` as the minimum number.
- If float, then `min_samples_split` is a fraction and
`ceil(min_samples_split * n_samples)` are the minimum
number of samples for each split.
.. versionchanged:: 0.18
Added float values for fractions.
min_samples_leaf : int or float, default=1
The minimum number of samples required to be at a leaf node.
A split point at any depth will only be considered if it leaves at
least ``min_samples_leaf`` training samples in each of the left and
right branches. This may have the effect of smoothing the model,
especially in regression.
- If int, then consider `min_samples_leaf` as the minimum number.
- If float, then `min_samples_leaf` is a fraction and
`ceil(min_samples_leaf * n_samples)` are the minimum
number of samples for each node.
.. versionchanged:: 0.18
Added float values for fractions.
min_weight_fraction_leaf : float, default=0.0
The minimum weighted fraction of the sum total of weights (of all
the input samples) required to be at a leaf node. Samples have
equal weight when sample_weight is not provided.
max_depth : int, default=3
The maximum depth of the individual regression estimators. The maximum
depth limits the number of nodes in the tree. Tune this parameter
for best performance; the best value depends on the interaction
of the input variables.
min_impurity_decrease : float, default=0.0
A node will be split if this split induces a decrease of the impurity
greater than or equal to this value.
The weighted impurity decrease equation is the following::
N_t / N * (impurity - N_t_R / N_t * right_impurity
- N_t_L / N_t * left_impurity)
where ``N`` is the total number of samples, ``N_t`` is the number of
samples at the current node, ``N_t_L`` is the number of samples in the
left child, and ``N_t_R`` is the number of samples in the right child.
``N``, ``N_t``, ``N_t_R`` and ``N_t_L`` all refer to the weighted sum,
if ``sample_weight`` is passed.
.. versionadded:: 0.19
init : estimator or 'zero', default=None
An estimator object that is used to compute the initial predictions.
``init`` has to provide :meth:`fit` and :meth:`predict_proba`. If
'zero', the initial raw predictions are set to zero. By default, a
``DummyEstimator`` predicting the classes priors is used.
random_state : int, RandomState instance or None, default=None
Controls the random seed given to each Tree estimator at each
boosting iteration.
In addition, it controls the random permutation of the features at
each split (see Notes for more details).
It also controls the random splitting of the training data to obtain a
validation set if `n_iter_no_change` is not None.
Pass an int for reproducible output across multiple function calls.
See :term:`Glossary <random_state>`.
max_features : {'auto', 'sqrt', 'log2'}, int or float, default=None
The number of features to consider when looking for the best split:
- If int, then consider `max_features` features at each split.
- If float, then `max_features` is a fraction and
`int(max_features * n_features)` features are considered at each
split.
- If 'auto', then `max_features=sqrt(n_features)`.
- If 'sqrt', then `max_features=sqrt(n_features)`.
- If 'log2', then `max_features=log2(n_features)`.
- If None, then `max_features=n_features`.
Choosing `max_features < n_features` leads to a reduction of variance
and an increase in bias.
Note: the search for a split does not stop until at least one
valid partition of the node samples is found, even if it requires to
effectively inspect more than ``max_features`` features.
verbose : int, default=0
Enable verbose output. If 1 then it prints progress and performance
once in a while (the more trees the lower the frequency). If greater
than 1 then it prints progress and performance for every tree.
max_leaf_nodes : int, default=None
Grow trees with ``max_leaf_nodes`` in best-first fashion.
Best nodes are defined as relative reduction in impurity.
If None then unlimited number of leaf nodes.
warm_start : bool, default=False
When set to ``True``, reuse the solution of the previous call to fit
and add more estimators to the ensemble, otherwise, just erase the
previous solution. See :term:`the Glossary <warm_start>`.
validation_fraction : float, default=0.1
The proportion of training data to set aside as validation set for
early stopping. Must be between 0 and 1.
Only used if ``n_iter_no_change`` is set to an integer.
.. versionadded:: 0.20
n_iter_no_change : int, default=None
``n_iter_no_change`` is used to decide if early stopping will be used
to terminate training when validation score is not improving. By
default it is set to None to disable early stopping. If set to a
number, it will set aside ``validation_fraction`` size of the training
data as validation and terminate training when validation score is not
improving in all of the previous ``n_iter_no_change`` numbers of
iterations. The split is stratified.
.. versionadded:: 0.20
tol : float, default=1e-4
Tolerance for the early stopping. When the loss is not improving
by at least tol for ``n_iter_no_change`` iterations (if set to a
number), the training stops.
.. versionadded:: 0.20
ccp_alpha : non-negative float, default=0.0
Complexity parameter used for Minimal Cost-Complexity Pruning. The
subtree with the largest cost complexity that is smaller than
``ccp_alpha`` will be chosen. By default, no pruning is performed. See
:ref:`minimal_cost_complexity_pruning` for details.
.. versionadded:: 0.22
Examples
--------
The following example shows how to fit a gradient boosting classifier with
100 decision stumps as weak learners.
>>> from sklearn.datasets import make_hastie_10_2
>>> from sklearn.ensemble import GradientBoostingClassifier
>>> X, y = make_hastie_10_2(random_state=0)
>>> X_train, X_test = X[:2000], X[2000:]
>>> y_train, y_test = y[:2000], y[2000:]
>>> clf = GradientBoostingClassifier(n_estimators=100, learning_rate=1.0,
... max_depth=1, random_state=0).fit(X_train, y_train)
>>> clf.score(X_test, y_test)
0.913...
File: c:\users\chjy1\anaconda3\lib\site-packages\sklearn\ensemble\_gb.py
Type: ABCMeta
Subclasses: