시작하며
오늘은 연휴 전 마지막 수업이었다. 의사결정나무에 이어 앙상블, 랜덤포레스트에 대한 내용을 배웠다.
의사결정나무 분류
의사결정나무 개요
- 분류와 회귀 문제에 널리 쓰이는 대표적인 알고리즘
- 각 노드에서 타겟의 불순도 계산
- 지니 계수, 엔트로피, 카이제곱 통계량 등을 사용
- 노드에서 타겟의 클래스가 완전하게 섞여 있을 때 불순도는 최대값
- 각 특성에 대해 분할 기준을 찾고, 분할 후 자식 노드에서의 불순도를 계산
- 부모 노드 → 자식 노드
- 불순도 감소량이 가장 큰 분할 기준을 선택
결정 트리의 장점
- 각 분할 기준을 바탕으로 IF-THEN 형태의 규칙을 추출
- 모델을 해석하고 실제 의사결정에 바로 활용하기 좋음
- 앙상블 알고리즘은 일반적으로 성능이 뛰어나지만 해석이 어려움
- 결정 트리는 해석이 용이해 현장에서 널리 사용 됨
- 비모수적 알고리즘으로 데이터 분포에 대한 사전 가정이 필요 없음
- 분할 기준으로 사용된 특성의 불순도 감소량을 집계
- 예측에 중요한 역할을 한 특성을 정량적으로 확인(특성중요도)
- 연속형 특성의 분포를 기준으로 분할 임계점을 자동 탐색
- 결측값을 하나의 범주로 간주하거나 대체 분할 기준을 사용
- Python에서는 결측 처리 필수
결정 트리의 단점
- 과적합 발생이 쉬움
- 모델 복잡 → 리프수 증가 → 관측 데이터 감소
과적합 방지
- 정지규칙 설정
- 나무가 무한히 성장하지 않도록 미리 제한하는 기준
- 자식 노드가 가져야 할 최소 관측값 개수 및 최대 깊이 등 포함
- 사후 가지치기
- 비용 복잡도 기준으로 불필요한 가지를 잘라내는 것
- 교차검증을 통해 성능이 우수한 비용 복잡도 파라미터 탐색
분류 나무 분할 기준
- 각 노드에서 가능한 모든 특성-입계값(분리 기준점) 조합을 평가(불순도 측정)
- 연속형 변수는 특정 임계값을 기준으로 분할
- 모든 고유값의 중간값 후보를 순차적으로 탐색
- 부모 노드에서 자식 노드의 불순도를 뺀 감소량이 가장 큰 분할 기준 선택
- 자식 노드의 불순도는 각 자식 노드의 불순도를 관측값 비율로 가중평균
결정 트리 알고리즘 종류
| 구분 | 불순도 | 특징 |
|---|---|---|
| CART | 지니 계수 사용 | 이진 분할 / 재귀적 |
| C5.0 | 엔트로피 사용 | 다중 분할 / 재귀적 |
| CHAID | 카이제곱 통계량 사용 | 다중 분할 / 범주형 특성만 처리 |
지니 계수
- 지니 계수는 0~0.5 범위의 값을 가짐
- 타겟이 한 클래스에 몰려 있을 때( 가 0 / 1) 지니 계수는 0
- 타겟의 클래스별 비율이 50:50일 때( 가 각각 0.5) 지니 계수는 0.5
- 이진 분류의 경우 지니 계수가 0일 때 불순도가 가장 낮고, 0.5일 때 가장 높음
- 자식 노드의 지니 계수를 각 자식 노드의 관측값 비율로 가중평균
- 지니 계수 감소량이 큰 분할 기준을 선택
엔트로피
- 엔트로피는 불확실성(무질서도)의 척도
- 이진 분류에서는 0~1가이의 값을 가짐
- 타겟이 한 클래스에 몰려 있을 때 엔트로피는 0
- 타겟의 클래스별 비율이 50:50일 때 엔트로피는 1
- 엔트로피가 1일 때 가장 무질서한 상태를 의미
- 엔트로피 감소량(정보 이득)이 큰 분할 기준을 선택
정보 이득비
- 다중 분할에서 자식 노드 개수가 늘어날수록 엔트로피가 감소하는 경향
- 정보 이득 기준 → 자식 노드가 많이지는 분할을 선호 → 분할의 내재 정보
- 내재 정보는 분할 자체가 얼마나 정보를 많이 소비했는지를 나타냄
- 자식 노드가 많아질수록 내재 정보가 커짐
- C5.0은 정보 이득비(정보 이득 / 내재 정보)를 분할 기준으로 사용
- 정보 이득비가 클수록 분할이 효율적
결정 트리의 특성 중요도
- 트리 모델을 학습하면 각 분할에 사용된 특성별 불순도 감소량을 계산할 수 있음
- 이를 모두 더하면 특성 중요도가 됨
- 분류 모델 : Gini / Entropy
- 회귀 모델 : 평균 제곱 오차(MSE)
- 각 특성별 불순도 감소량을 전체 특성의 불순도 감소량의 합계로 나누어 정규화
- 모든 특성 중요도의 합은 1
- 각 특성이 총불순도 감소량에 얼마나 기여했는지 파악 가능
- 과적합된 트리는 특정 특성에 지나치게 높은 중요도를 부여할 수 있음
- 한 번에 하나의 특성으로 분할 → 특성 간 상호작용 효과를 과소평가할 수 있음
가지치기 전 분류 모델 학습
from sklearn.tree import DecisionTreeClassifiercriterion: 분할 품질 측정 기준 gini, entropy, log_loss- 기본값 : gini
max_depth: 트리의 최대 깊이 설정- 기본값 : None
min_samples_split: 부모 노드를 분할하기 위한 최소 샘플 개수를 설정- 기본값 : 2
min_samples_leaf: 자식 노드로 분할되기 위한 최소 샘플 개수를 설정- 기본값 : 1
max_features: 각 노드를 분할할 때 후보로 고려할 특성 개수를 설정- 기본값 : None
ccp_alpha: 가지치기 비용 복잡도 파라미터 → 사후 가지치기를 수행할 때 사용
model_full = DecisionTreeClassifier(min_samples_split=100, random_state=0)
model_full.fit(X=X_train, y=y_train)model_full.score(X=X_train, y=y_train)
# 0.8445157526254375
model_full.score(X=X_valid, y=y_valid)
# 0.791156462585034- 모델의 특성 중요도 확인
pd.Series(data=model_full.feature_importances_, index=model_full.feature_names_in_).sort_values(ascending=False)
# alcohol 0.532137
# volatile acidity 0.101741
# pH 0.067792
# residual sugar 0.067128
# chlorides 0.058272
# density 0.055429
# sulphates 0.049044
# total sulfur dioxide 0.034607
# fixed acidity 0.033050
# citric acid 0.000800
# dtype: float64hds.plot.feature_importance(model_full)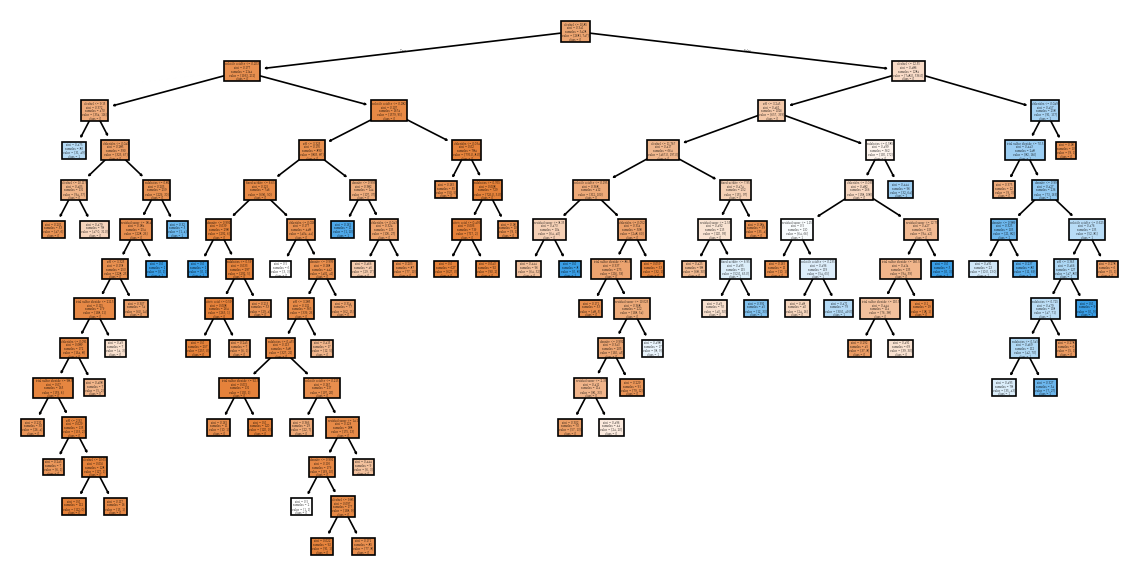
- 터미널 노드 개수 확인
model_full.get_n_leaves()
# np.int64(62)- 최대 깊이 조회
model_full.get_depth()
# 13트리 모델 시각화
from sklearn.tree import plot_treeplt.figure(figsize=(12, 6))
plot_tree(model_full, feature_names=X_train.columns, class_names=model_full.classes_.astype(str), filled=True)
plt.show()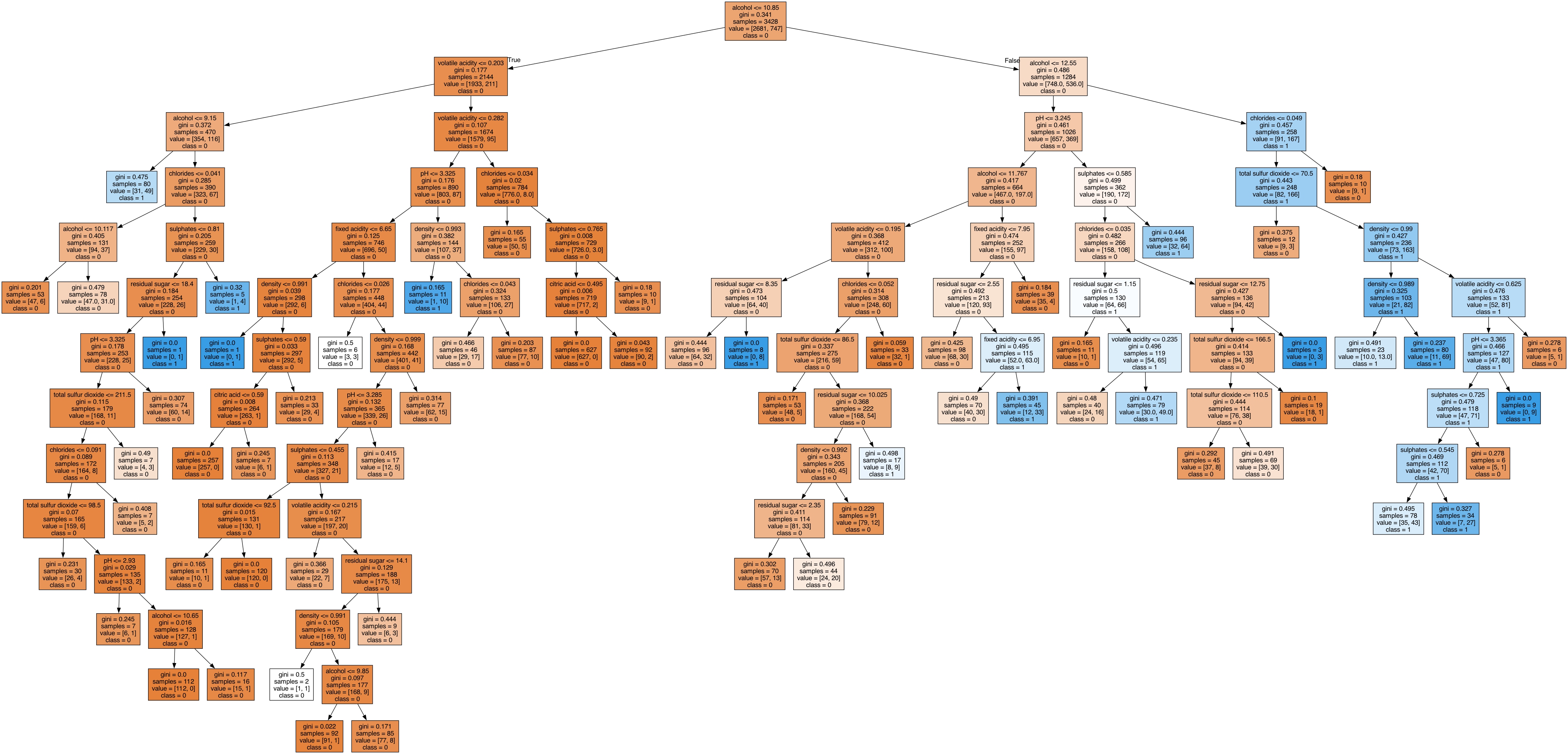
- 가독성 좋게 png 파일로 저장
hds.plot.tree(model=model_full, fileName='dtc_full')from IPython.display import Image
Image('dtc_full.png')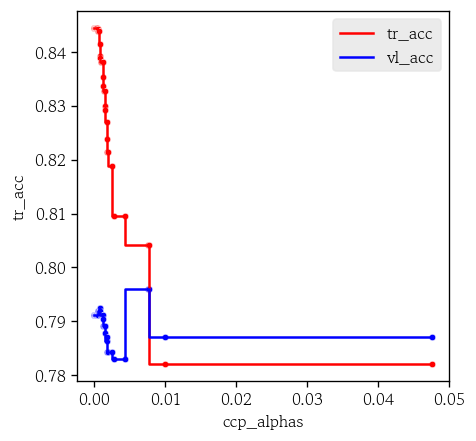
사후 가지치기
- 학습을 완료하면 과적합 방지를 위해 사후 가지치기 필요성 검토
- 비용 복잡도를 이용 → 복잡도 가지치기(CCP)
- : 비용 복잡도 함수
- : 트리 모델 T의 성능 손실 지표
- : 비용 복잡도 파라미터로 0보다 큰 실수
- : 트리 모델의 리프 노드 개수
- 리프 노드 개수가 증가하면 복잡도가 높아짐
- 리프 노드의 개수 줄어듦 → 노드에 포함되는 관측값 개수 증가
사후 가지치기 경로 확인
path = model_full.cost_complexity_pruning_path(X_train, y_train)
path = pd.DataFrame(path)
path.head()
# ccp_alphas impurities
# 0 0.000000 0.211743
# 1 0.000022 0.211765
# 2 0.000032 0.211797
# 3 0.000049 0.211845
# 4 0.000054 0.211900최적의 비용 복잡도 파라미터() 탐색
from sklearn.base import clone- 가지치기 전 모델을 복제
def clone_tree(alpha):
model = clone(model_full)
model.set_params(ccp_alpha=alpha)
model.fit(X_train, y_train)
return model- 다양한 비용 복잡도 파라미터를 설정한 복제 모델을 원소로 갖는 리스트 생성
trees = [clone_tree(alpha) for alpha in path['ccp_alphas']]path['leaves'] = [tree.get_n_leaves() for tree in trees]
path['tr_acc'] = [tree.score(X_train, y_train) for tree in trees]
path['vl_acc'] = [tree.score(X_valid, y_valid) for tree in trees]
path.head()
# ccp_alphas impurities leaves tr_acc vl_acc
# 0 0.000000 0.211743 62 0.844516 0.791156
# 1 0.000022 0.211765 61 0.844516 0.791156
# 2 0.000032 0.211797 60 0.844516 0.791156
# 3 0.000049 0.211845 59 0.844516 0.791156
# 4 0.000054 0.211900 58 0.844516 0.791156- 시각화
hds.plot.step(data=path, x='ccp_alphas', y='tr_acc', color='red')
hds.plot.step(data=path, x='ccp_alphas', y='vl_acc', color='blue')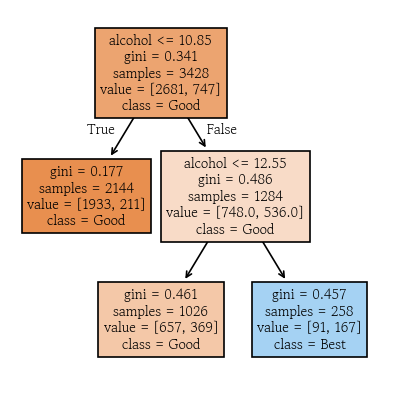
- 검증셋 정확도가 최댓값인 인덱스 확인
np.argmax()는 최댓값이 여러 개 있으면 맨 처음 인덱스를 반환
np.argmax(path['vl_acc'])
# np.int64(41)- 오름차순 정렬한 원소의 인덱스 생성
indices = np.argsort(path['vl_acc'])
indices.iloc[-1]
# np.int64(42)- 정확도가 최댓값일 때의 alpha를 best_alpha에 할당
best_alpha = path['ccp_alphas'][indices.iloc[-1]]
# np.float64(0.0077337653548115864)가지치기 후 분류 모델 학습
- 새로운 모델 복제
model_prun = clone(model_full)- best_alpha 설정 후 모델 학습
model_prun.set_params(ccp_alpha=best_alpha)
model_prun.fit(X_train, y_train)- 정확도 확인
model_prun.score(X_train, y_train)
# 0.8042590431738623
model_prun.score(X_valid, y_valid)
# 0.7959183673469388- 깊이 및 리프 개수 확인
model_prun.get_depth()
# 2
model_prun.get_n_leaves()
# np.int64(3)- 시각화
plot_tree(model_prun, feature_names=X_train.columns, class_names=['Good', 'Best'], filled=True)
plt.show()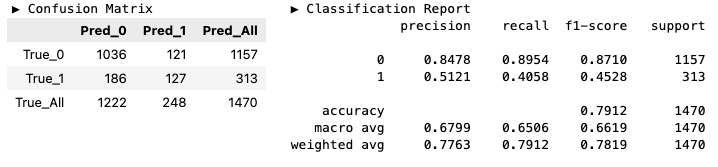
분류 모델 성능 평가
y_pred_full = model_full.predict(X_valid)
y_pred_prun = model_prun.predict(X_valid)hds.stat.clfmetrics(y_true=y_valid, y_pred=y_pred_full)
hds.stat.clfmetrics(y_true=y_valid, y_pred=y_pred_prun)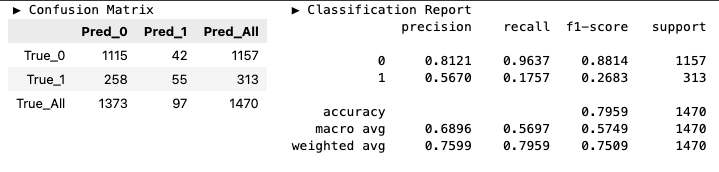
ROC / PR 곡선
- 예측 활률 생성
y_prob_full = model_full.predict_proba(X_valid)
y_prob_prun = model_prun.predict_proba(X_valid)- ROC 곡선
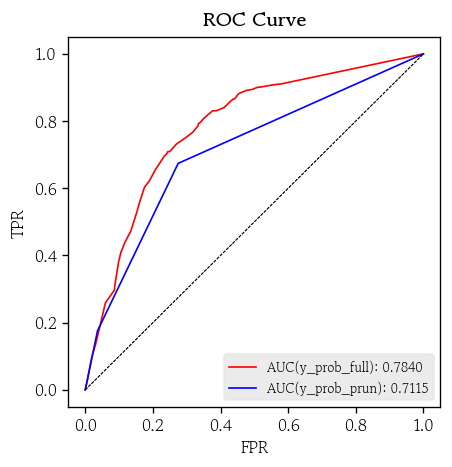
hds.plot.roc_curve(y_true=y_valid, y_prob=y_prob_full, color='red')
hds.plot.roc_curve(y_true=y_valid, y_prob=y_prob_prun, color='blue')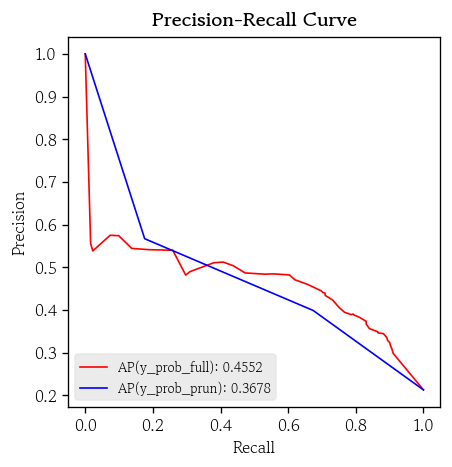
- PR 곡선
hds.plot.pr_curve(y_true=y_valid, y_prob=y_prob_full, color='red')
hds.plot.pr_curve(y_true=y_valid, y_prob=y_prob_prun, color='blue')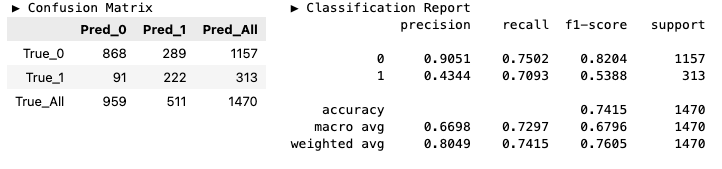
불균형 데이터셋에 대한 처리 방법
타겟 벡터의 실제값 상대도수로 분류 기준점 설정
y_valid.value_counts(normalize=True)
# grade
# 0 0.787075
# 1 0.212925
# # Name: proportion, dtype: float64cutoff = 0.212925
y_pred_best_0 = np.where(y_prob_full[:, 1] >= cutoff, 1, 0)
hds.stat.clfmetrics(y_true=y_valid, y_pred=y_pred_best_0)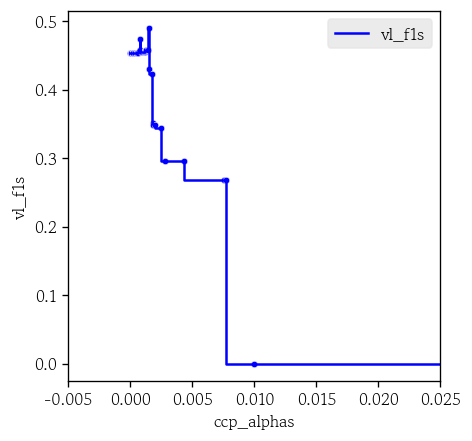
검증셋 정확도 대신 F1 점수 기준으로 가지치기
from sklearn.metrics import f1_score- f1 스코어 계산 함수
def valid_f1_score(tree):
y_pred = tree.predict(X=X_valid)
score = f1_score(y_true=y_valid, y_pred=y_pred)
return score- f1스코어 값 컬럼 생성
path['vl_f1s'] = [valid_f1_score(tree) for tree in trees]hds.plot.step(data=path, x='ccp_alphas', y='vl_f1s')
plt.xlim(-0.005, 0.025)
plt.show()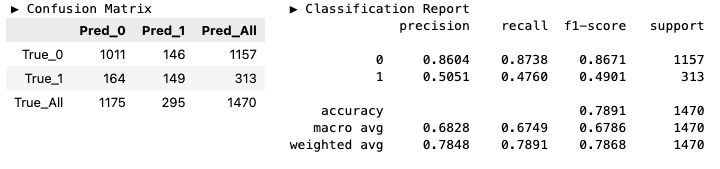
index = np.argsort(path['vl_f1s'])
best_alpha_f1s = path['ccp_alphas'][index.iloc[-1]]
# np.float64(0.0014843053552403666)model_best_1 = clone(model_full)
model_best_1.set_params(ccp_alpha=best_alpha_f1s)
model_best_1.fit(X_train, y_train)
y_pred_best_1 = model_best_1.predict(X_valid)
hds.stat.clfmetrics(y_true=y_valid, y_pred=y_pred_best_1)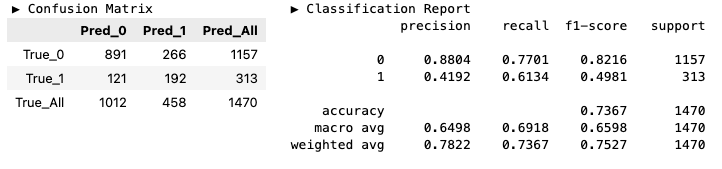
SMOTE를 활용한 데이터셋 균형화
from imblearn.over_sampling import SMOTEsmote = SMOTE(k_neighbors=5, random_state=0)
X_bal, y_bal = smote.fit_resample(X_train, y_train)
model_best_2 = DecisionTreeClassifier(min_samples_split=100, random_state=0)
model_best_2.fit(X_bal, y_bal)model_best_2.score(X_bal, y_bal)
# 0.8508019395747856
model_best_2.score(X_valid, y_valid)
# 0.736734693877551y_pred_best_2 = model_best_2.predict(X_valid)
hds.stat.clfmetrics(y_true=y_valid, y_pred=y_pred_best_2)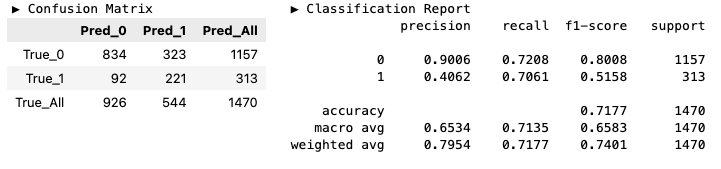
class_weight 매개변수에 'balanced' 지정하여 학습
model_best_3 = clone(model_full)
model_best_3.set_params(class_weight='balanced')
model_best_3.fit(X_train, y_train)model_best_3.score(X_train, y_train)
# 0.793757292882147
model_best_3.score(X_valid, y_valid)
# 0.717687074829932y_pred_best_3 = model_best_3.predict(X_valid)
hds.stat.clfmetrics(y_true=y_valid, y_pred=y_pred_best_3)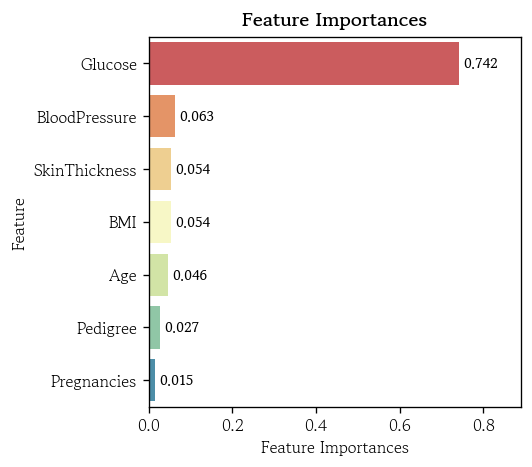
의사결정나무 회귀
회귀 나무 분할 기준
- 분할 기준으로 평균 제곱 오차(MSE)의 감소량을 사용
- MSE 감소량을 최대화하는 분할을 선택
- 타겟의 변동성(분산)이 작은 관측값을 같은 자식 노드에 모이도록 함
- 범주형이거나 특성과 타겟 간의 관계가 비선형일 때 선형 회귀보다 더욱 유연하고 우수
가지치기 전 회귀 모델 학습
from sklearn.tree import DecisionTreeRegressormodel_full = DecisionTreeRegressor(min_samples_split=30, random_state=0)
model_full.fit(X_train, y_train)model_full.score(X_train, y_train)
# 0.716395830725133
model_full.score(X_valid, y_valid)
# 0.35409551601258393- 특성 중요도 확인
pd.Series(data=model_full.feature_importances_, index=model_full.feature_names_in_).sort_values(ascending=False)
# Glucose 0.741621
# BloodPressure 0.062569
# SkinThickness 0.054211
# BMI 0.053741
# Age 0.045612
# Pedigree 0.026888
# Pregnancies 0.015359
# dtype: float64hds.plot.feature_importance(model_full)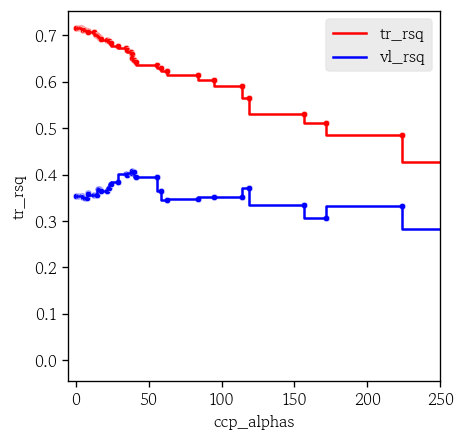
사후 가지치기 경로 확인
path = model_full.cost_complexity_pruning_path(X_train, y_train)
path = pd.DataFrame(path)
path.head()
# ccp_alphas impurities
# 0 0.000000 2562.532263
# 1 2.634336 2565.166599
# 2 3.450466 2568.617064
# 3 3.849355 2572.466420
# 4 4.250502 2580.967423최적의 비용 복잡도 파라미터 탐색
from sklearn.base import clonedef clone_tree(alpha):
model = clone(model_full)
model.set_params(ccp_alpha=alpha)
model.fit(X_train, y_train)
return modeltrees = [clone_tree(alpha) for alpha in path['ccp_alphas']]path['leaves'] = [tree.get_n_leaves() for tree in trees]
path['tr_rsq'] = [tree.score(X_train, y_train) for tree in trees]
path['vl_rsq'] = [tree.score(X_valid, y_valid) for tree in trees]hds.plot.step(data=path, x='ccp_alphas', y='tr_rsq', color='red')
hds.plot.step(data=path, x='ccp_alphas', y='vl_rsq', color='blue')
plt.xlim(-5, 250)
plt.show()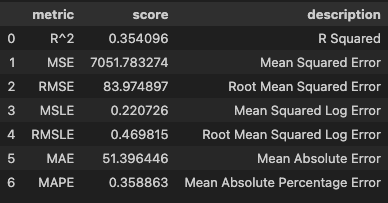
index = np.argsort(path['vl_rsq'])
best_alpha = path['ccp_alphas'][index.iloc[-1]]
# np.float64(38.711664229416954)model_prun = clone(model_full)
model_prun.set_params(ccp_alpha=best_alpha)
model_prun.fit(X_train, y_train)y_pred_full = model_full.predict(X_valid)
y_pred_prun = model_prun.predict(X_valid)hds.stat.regmetrics(y_true=y_valid, y_pred=y_pred_full)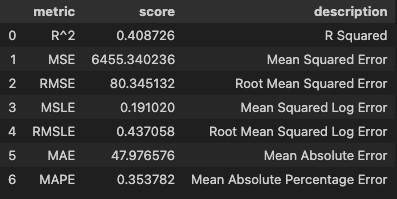
hds.stat.regmetrics(y_true=y_valid, y_pred=y_pred_prun)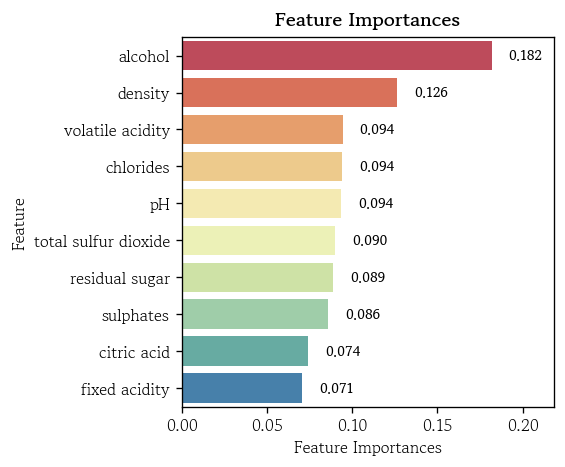
Ensemble
앙상블 기법 개요
- 여러 개의 단일 모델을 동시에 학습 → 예측 결과를 종합 → 전체 성능을 향상
- 한계를 극복하여 일반화 성능을 높일 수 있음
- 여러 모델을 결합하기 때문에 기여도 파악 및 해석이 복잡함
보팅
- 서로 다른 알고리즘으로 학습한 여러 개 모델의 예측값을 결합하는 방식
- 하드 보팅
- 예측값 다수결
- 소프트 보팅
- 예측 확률 평균
배깅
- Bootstrapping과 Aggregating의 합성어
- 훈련셋에서 동일한 크기의 샘플을 복원 추출로 뽑아 각각 독립적인 모델 학습
- 예측값을 종합하여 최종 결과를 얻는 기법
- 원본 훈련셋에서 중복을 허용하여 샘플 추출
- 표본 평균의 기댓값은 모평균과 같고 표본 평균의 분산은 모분산을 샘플크기로 나눈 값이 됨
- 단일 결정 트리는 과적합에 최약하지만 수백~수천개의 트리를 학습하여 일반화 성능 향상
부스팅
- 순차적으로 이전 모델이 잘못 분류한 관측값에 가중치를 높여 다음 모델이 더 집중해서 학습하도록 함
- 약한 학습기(작은 나무)를 순차적으로 학습
- 각 분류기의 성능에 따라 오분류율에 반비례하는 가중치를 부여
- 모든 분류기를 가중합하여 강한 분류기로 결합
- 하이퍼파라미터를 잘못 설정하면 과적합이 발생하기 쉬움
데이터셋에 따른 앙상블 기법 추천
- 랜덤 포레스트
- 노이즈가 많거나 결측값이 있는 데이터
- 안정적인 성능이 필요
- 하이퍼파라미터 튜닝이 단순
- 과적합 방지 효과가 강렬(평균 → 평균)
- 병렬 학습을 활용해 학습 시간을 단축
- 모델의 복잡도가 너무 높지 않아도 잘 작동
- 그레디언트 부스팅
- 예측 정확도가 최우선
- 복잡한 비선형 상호작용을 잡아내는 작업
- 작은 성능 향상이라도 중요한 업무
- 하이퍼파라미터를 세밀하게 튜닝해야 효과 극대화
랜덤 포레스트 분류
- 원본 훈련셋에서 복원추출 방식으로 서로 다른 크기의 샘플을 n번 추출
- 독립적인 n개의 결정 트리를 학습
- 독립적 : 각 트리가 서로의 결과나 구조에 영향을 주지 않는 것
- 새로운 관측값을 학습된 n개의 트리에 입력
- n개의 예측을 얻음
- 다수결 방식 또는 평균을 계산하여 최종 예측값 반환
랜덤 포레스트 개요
- 수백~수천 개의 트리를 학습해 각각의 결과를 결합하는 앙상블 기법
- 다수의 트리가 협력하여 과적합을 줄이고 일반화 성능을 높임
- 개별 트리를 과적합시켜 편향을 최소화 → 앙상블로 분산을 줄임
랜덤 포레스트의 특징
- 훈련셋에서 부트스트랩 샘플을 서브셋으로 사용
- 똑같이 중요한 몇몇 특성에 의존하지 않게 됨
- 다양한 관점을 반영
- 학습할 때 사용하지 않은 Out-of-bag 샘플을 이용해 성능을 추정
- OOB 오분류율을 최소화하는 하이퍼파라미터 조합을 찾을 수 있음
- 그리드 서치, 랜덤 서치, 베이지안 최적화 등을 사용하여 교차 검증 성능이 좋은 조합을 탐색
랜덤 포레스트의 장점
- 여러 개의 트리를 독립적으로 학습시켜 과적합 위험을 줄임
- 다양한 패턴을 효과적으로 포착
- 각 트리는 서로 다른 샘플과 특성 서브셋으로 학습
- 노이즈에 강한 안정적인 예측 성능을 제공
- 특성 중요도를 제공 → 중요 특성을 직관적으로 확인 가능
- 학습 속도를 크게 향상
랜덤 포레스트의 단점
- 모델 구조가 복잡한 블랙박스 형태가 되어 내부 작동 원리를 파악하기 어려움
- 모든 트리를 거쳐 앙상블 투표를 수행
- 실시간 예측에는 적합하지 않을 수 있음
랜덤 포레스트의 특성 중요도
- 다수의 결정 트리를 앙상블 하여 학습
- 각 트리에서 계산된 정규화된 특성 중요도 합계를 트리 개수로 나누어 계산
OOB 정확도 추정
- 훈련셋을 부트스트랩 방식으로 샘플링
- 각 트리 학습에 사용되지 않은 관측값은 전체의 약 36%
- 데이터가 부족할 때 별도의 검증셋 없어도 OOB 샘플로 모델 성능을 평가
- 클래스 불균형이 심한 경우 정확도가 다소 편향되거나 불안정할 수 있음
랜덤 포레스트 분류 모델 학습
from sklearn.ensemble import RandomForestClassifiermodel = RandomForestClassifier(oob_score=True, random_state=0)
model.fit(X_train, y_train)model.score(X_train, y_train)
# 1.0
model.score(X_valid, y_valid)
# 0.8653061224489796pd.Series(data=model.feature_importances_, index=model.feature_names_in_).sort_values(ascending=False)
# alcohol 0.181730
# density 0.126302
# volatile acidity 0.094409
# chlorides 0.094020
# pH 0.093747
# total sulfur dioxide 0.090092
# residual sugar 0.088768
# sulphates 0.085994
# citric acid 0.074391
# fixed acidity 0.070548
# dtype: float64hds.plot.feature_importance(model)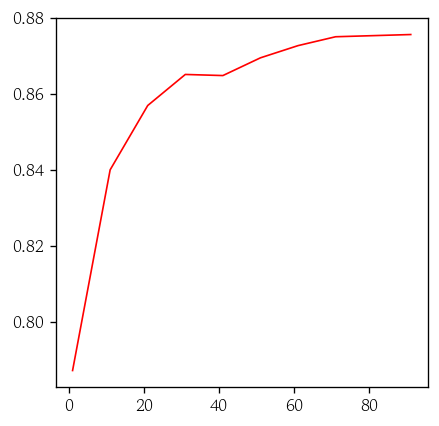
OOB 정확도 시각화
model.oob_score_
# 0.8771878646441074- OOB 정확도 반환 함수
def oob_score(ntree):
model.set_params(n_estimators=ntree)
model.fit(X_train, y_train)
return model.oob_score_- 트리 개수 범위 설정 및 정확도를 원소로 갖는 리스트 생성
ntrees = range(1, 101, 10)
oob_acc = [oob_score(ntree) for ntree in ntrees]sns.lineplot(x=ntrees, y=oob_acc, color='red', linewidth=1)
plt.show()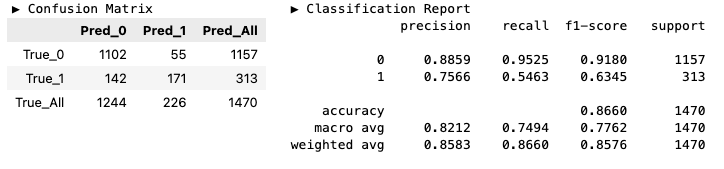
분류 모델 성능 평가
- 예측값 생성
y_pred = model.predict(X_valid)hds.stat.clfmetrics(y_true=y_valid, y_pred=y_pred)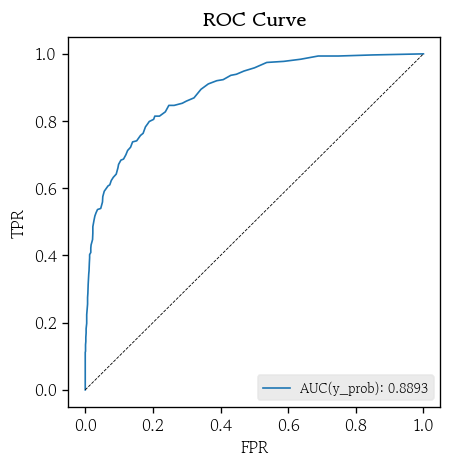
곡선 시각화
y_prob = model.predict_proba(X_valid)hds.plot.roc_curve(y_true=y_valid, y_prob=y_prob)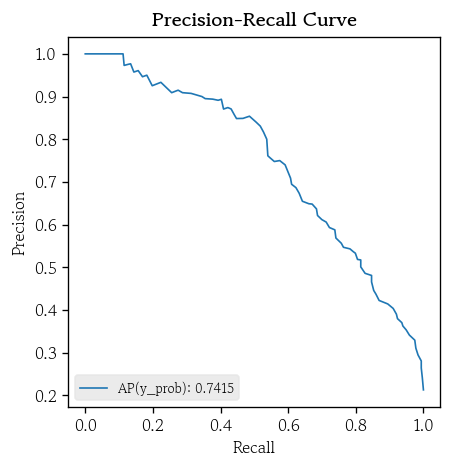
hds.plot.pr_curve(y_true=y_valid, y_prob=y_prob)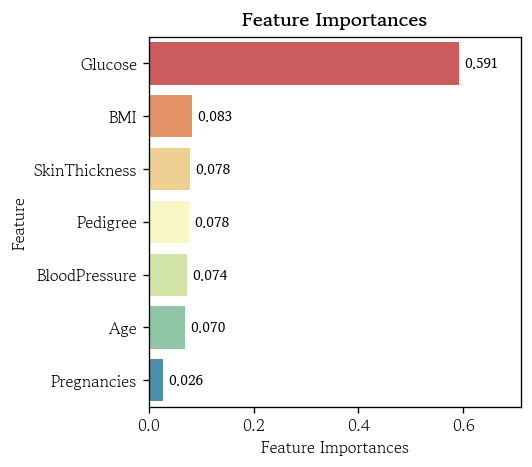
랜덤 포레스트 회귀
랜덤 포레스트 회귀 개요
- 여러 개의 회귀 나무를 학습하고, 얻은 예측값의 평균을 계산하여 최종 예측값으로 사용
- 각 회귀 나무의 예측값은 리프 노드의 평균값 → 평균값의 기대값을 제공
- 편향과 분산을 동시에 줄일 수 있음
- 일반화 성능 향상
랜덤 포레스트 회귀 모델 학습
from sklearn.ensemble import RandomForestRegressormodel = RandomForestRegressor(oob_score=True, random_state=0)
model.fit(X_train, y_train)model.score(X_train, y_train)
# 0.9202492412051448
model.score(X_valid, y_valid)
# 0.43848038611513096- 특성 중요도 확인
pd.Series(data=model.feature_importances_, index=model.feature_names_in_).sort_values(ascending=False)
# Glucose 0.591254
# BMI 0.082809
# SkinThickness 0.078039
# Pedigree 0.077711
# BloodPressure 0.073544
# Age 0.070146
# Pregnancies 0.026497
# dtype: float64hds.plot.feature_importance(model)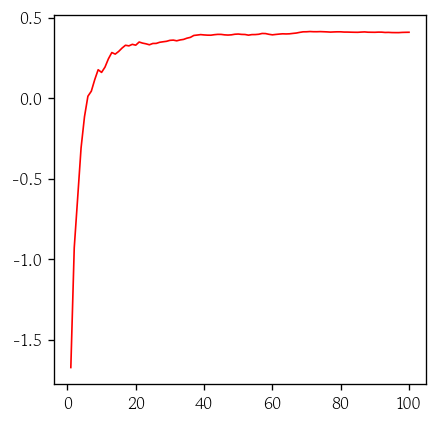
OOB 결정계수 시각화
def oob_score(ntree):
model.set_params(n_estimators=ntree)
model.fit(X_train, y_train)
return model.oob_score_ntrees = range(1, 101)
oob_rsq = [oob_score(ntree) for ntree in ntrees]sns.lineplot(x=ntrees, y=oob_rsq, color='red', linewidth=1)회귀 모델 성능 평가
- 예측값 생성
y_pred = model.predict(X_valid)hds.stat.regmetrics(y_true=y_valid, y_pred=y_pred)마치며
내일부터는 연휴가 시작되니까 그동안 부족했던 내용들과 찾아보려 했던 내용들을 공부해봐야겠다.
